Computes a one-sided upper tolerance interval for censored data assuming log-normal, gamma and normal distributions.
cenTolInt(
x.var,
cens.var,
conf = 0.95,
cover = 0.9,
method.fit = "mle",
printstat = TRUE
)Arguments
- x.var
The column of x (response variable) values plus detection limits
- cens.var
The column of indicators, where 1 (or
TRUE) indicates a detection limit in they.varcolumn, and 0 (orFALSE) indicates a detected value iny.var.- conf
Confidence coefficient of the interval, 0.95 (default).
- cover
Coverage, the percentile probability above which the tolerance interval is computed. The default is 90, so a tolerance interval will be computed above the 90th percentile of the data.
- method.fit
The method used to compute the parameters of the distribution. The default is maximum likelihood (
“mle”). The alternative is robust ROS (“rROS”). See Details.- printstat
Logical
TRUE/FALSEoption of whether to print the resulting statistics in the console window, or not. Default isTRUE.
Value
Prints and returns the percentile (cover), upper tolerance limit (conf) and BIC of fit for lognormal, normal and approximated gamma distributions. Plots empirical and theoretical CDFs with BIC values in the legend.
Details
Computes upper one-sided tolerance intervals for three distributions. This is a front-end to the individual functions from the EnvStats package. By default all three are computed using maximum likelihood estimation (mle); robust ROS is available as an alternate method for all three distributions. The gamma distribution for censored data uses the Wilson-Hilferty approximation (normal distribution on cube roots of data). For more info on the relative merits of robust ROS versus mle, see Helsel (2011) and Millard (2013).
References
Helsel, D.R., 2011. Statistics for censored environmental data using Minitab and R, 2nd ed. John Wiley & Sons, USA, N.J.
Millard, S.P., 2013. EnvStats: An R Package for Environmental Statistics. Springer-Verlag, New York.
Krishnamoorthy, K., Mathew, T., Mukherjee, S., 2008. Normal-Based Methods for a Gamma Distribution, Technometrics, 50, 69-78.
Examples
data(PbHeron)
# Default
cenTolInt(PbHeron$Liver,PbHeron$LiverCen)
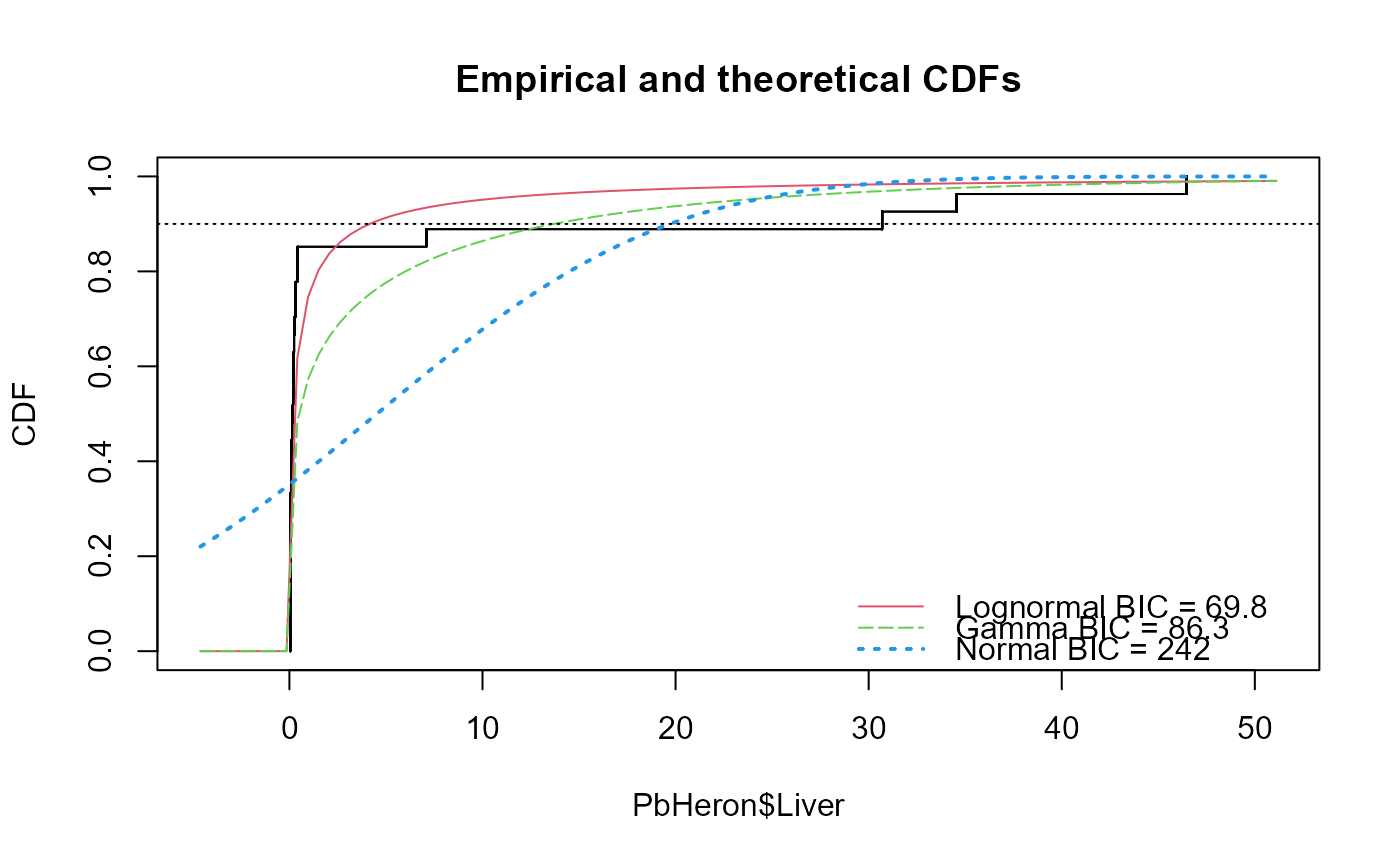 #> Distribution 90th Pctl 95% UTL BIC Method
#> 1 Lognormal 4.156275 14.48148 69.80314 mle
#> 2 Gamma 9.080911 18.22023 86.31145 mle
#> 3 Normal 19.957630 26.91080 242.48001 mle
# User defined conficence interval
cenTolInt(PbHeron$Liver,PbHeron$LiverCen,conf=0.75)
#> Distribution 90th Pctl 75% UTL BIC Method
#> 1 Lognormal 4.156275 6.743991 69.80314 mle
#> 2 Gamma 9.080911 12.130005 86.31145 mle
#> 3 Normal 19.957630 22.653852 242.48001 mle
# User defined percentile
cenTolInt(PbHeron$Liver,PbHeron$LiverCen,cover=0.5)
#> Distribution 90th Pctl 95% UTL BIC Method
#> 1 Lognormal 4.156275 14.48148 69.80314 mle
#> 2 Gamma 9.080911 18.22023 86.31145 mle
#> 3 Normal 19.957630 26.91080 242.48001 mle
# User defined conficence interval
cenTolInt(PbHeron$Liver,PbHeron$LiverCen,conf=0.75)
#> Distribution 90th Pctl 75% UTL BIC Method
#> 1 Lognormal 4.156275 6.743991 69.80314 mle
#> 2 Gamma 9.080911 12.130005 86.31145 mle
#> 3 Normal 19.957630 22.653852 242.48001 mle
# User defined percentile
cenTolInt(PbHeron$Liver,PbHeron$LiverCen,cover=0.5)
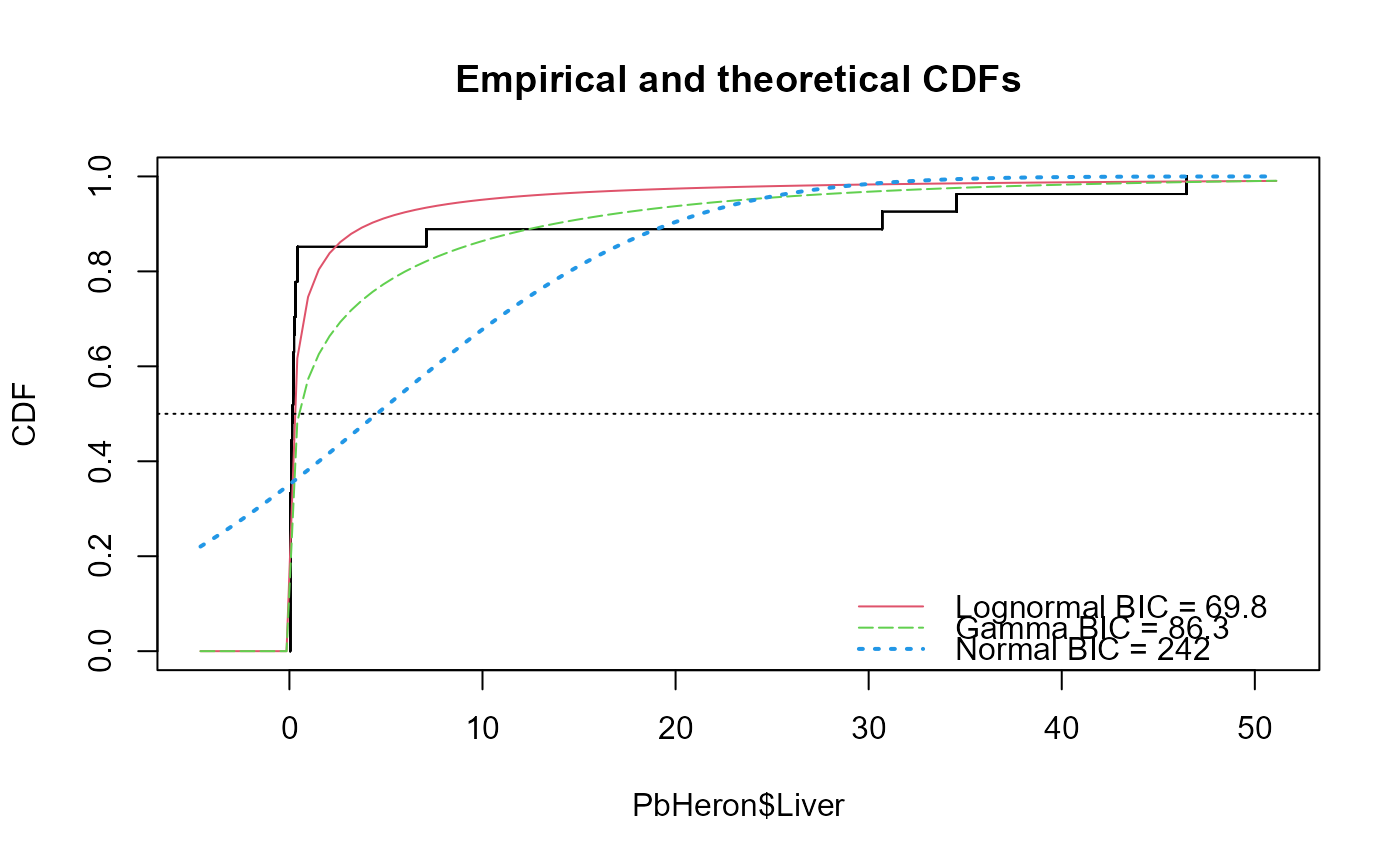 #> Distribution 50th Pctl 95% UTL BIC Method
#> 1 Lognormal 0.2029654 0.4398284 69.80314 mle
#> 2 Gamma 0.4526509 1.3511311 86.31145 mle
#> 3 Normal 3.1389176 7.4467275 242.48001 mle
# inputs outside acceptable ranges
# Will result in errors/warnings
# cenTolInt(PbHeron$Liver,PbHeron$LiverCen,cover=1.25)
# cenTolInt(PbHeron$Liver,PbHeron$LiverCen,conf=1.1)
# cenTolInt(PbHeron$Liver,PbHeron$LiverCen,method.fit="ROS")
#> Distribution 50th Pctl 95% UTL BIC Method
#> 1 Lognormal 0.2029654 0.4398284 69.80314 mle
#> 2 Gamma 0.4526509 1.3511311 86.31145 mle
#> 3 Normal 3.1389176 7.4467275 242.48001 mle
# inputs outside acceptable ranges
# Will result in errors/warnings
# cenTolInt(PbHeron$Liver,PbHeron$LiverCen,cover=1.25)
# cenTolInt(PbHeron$Liver,PbHeron$LiverCen,conf=1.1)
# cenTolInt(PbHeron$Liver,PbHeron$LiverCen,method.fit="ROS")